enzymes:
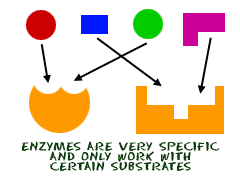
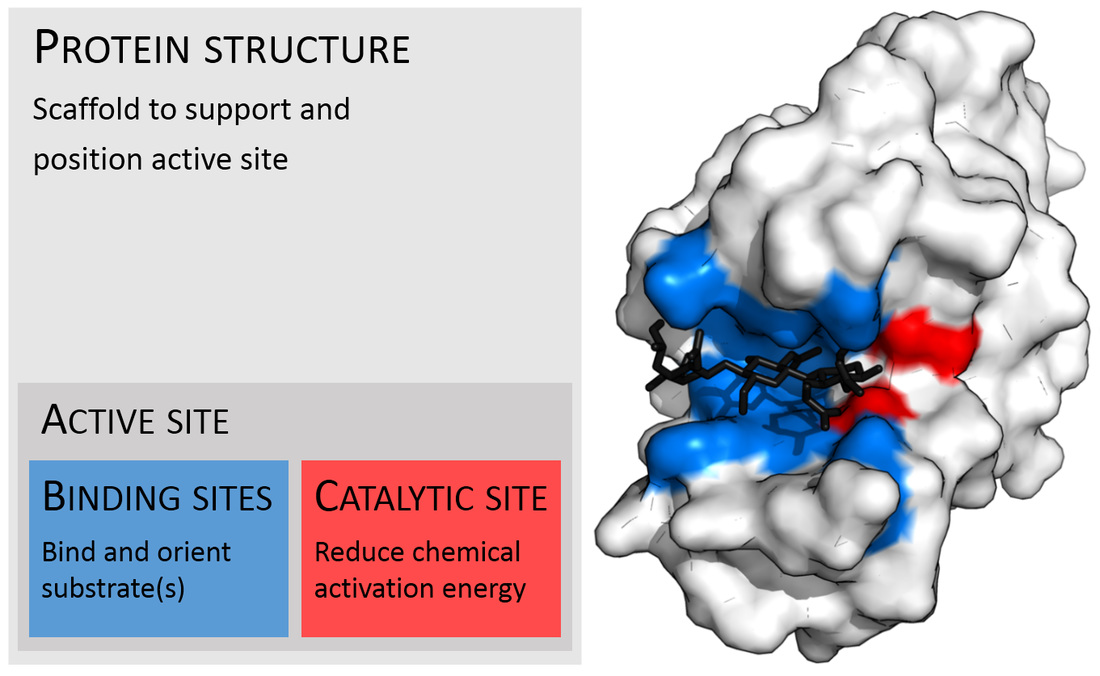
Enzymes are macromolecules (large, 3D or 4D protein molecules) that catalyze, or kick off, or speed up biochemical reactions. Substrates bind to and act on the enzyme, which converts them into products. Enzymes increase the rate of a biochemical reaction. They are very very specific, like a lock and key. They break down larger molecules into smaller ones needed to sustain life. Some enzymes require coenzymes or cofactors in order to kick off a reaction.
Many antimicrobials act on enzymes or block them from doing their job, so the bacteria die.
Many antimicrobials act on enzymes or block them from doing their job, so the bacteria die.
penicillin-binding protein and destruction of this protein by penicillin and antibiotic resistance
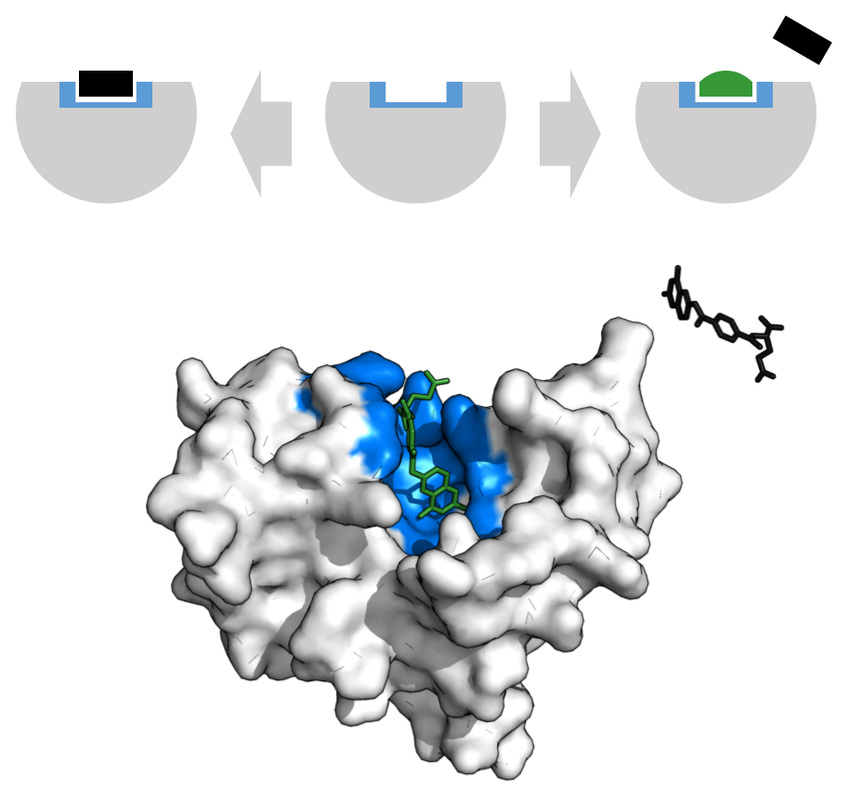
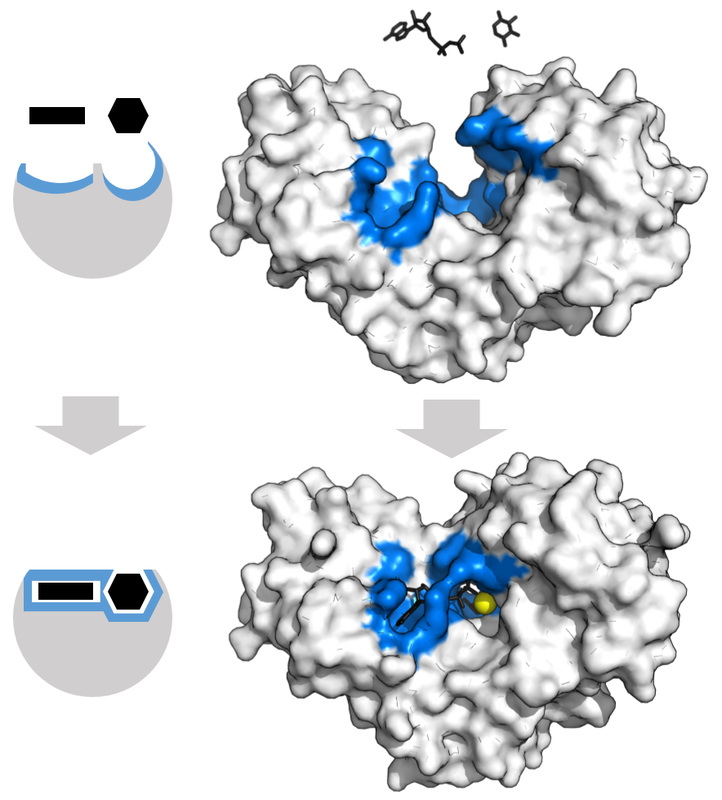
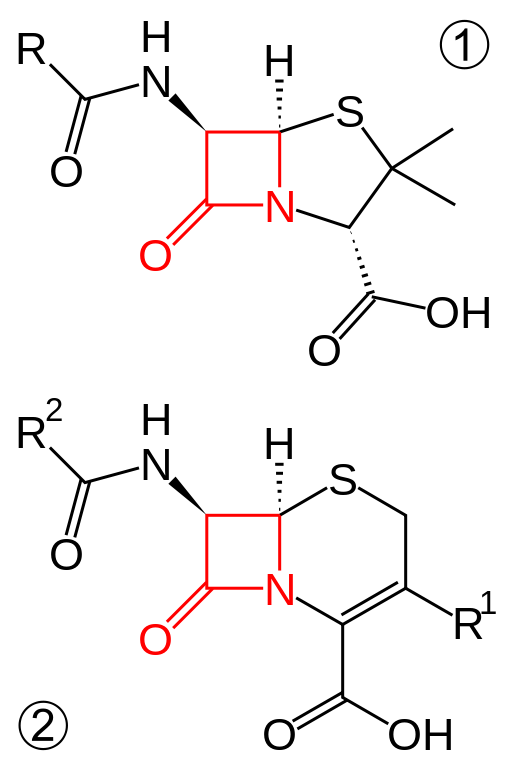
Penicillin Binding Proteins (PBPs) are proteins (enzymes) present in the cell walls of some bacteria. They catalyze the cross-linking of peptidoglycan in the bacterial cell walls. They have an internal structure similar to that of Penicillin and other Beta-lactam antibiotics, therefore the antibiotics work by binding this structure, thereby inhibiting cell wall synthesis. PBPs are critical for cell wall synthesis, and without them, the cell wall becomes disrupted, loses membrane potential, loses it selective membrane permeability, and the cell dies and lyses.
There are a large number of these proteins, and each bacterial prototype may have several of them. PBPs are involved in the last stages of peptidoglycan synthesis and the cross-linking of the cell wall for its shape and structure. This is essential for bacterial growth, survival, and cellular division and reproduction. Destruction of PBPs by the Beta-lactam antibiotics results in cell wall irregularities, elongation (can be seen in the Gram stain of patients on these antibiotics), and cell death. When the Beta-lactam antibiotic binds to this protein, the Beta-lactam amide bond is broken. This results in the formation of a covalent bond at the PBP's active site with the catalytic serine residue, and this bond is irreversible, thereby inactivating the protein permanently.
In cases of antibiotic resistance to Beta-lactams, such as in Methicillin-Resistant Staphylococcus Aureus, its PBP2A protein has a low affinity for penicillin and methicillin and the Beta-lactams, so the bond is not tight. This permits the protein to continue synthesizing the cell wall in the presence of the antibiotic and transpeptidase enzyme. This prevents the inhibiting of cell wall synthesis and renders the bacterium resistant to the drug(s). The presence of the PBP2A protein, and the presence of the mecA gene which encodes for it, enable this bacterium a resistant organism.
There are several rapid microbiology tests that detect the presence of the PBP2A protein or the mecA gene. These include the:
1) PBP2A rapid test, a quick 5 minute screen that detects the presence of this protein and presumptive MRSA
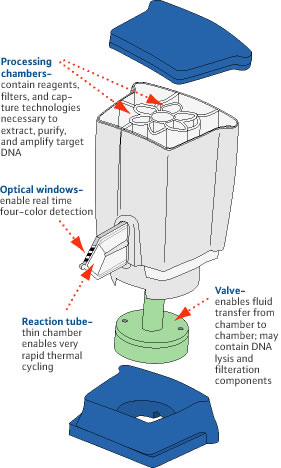
2) MRSA PCR (GenEXPERT), a one hour test that detects the presence of the mecA gene
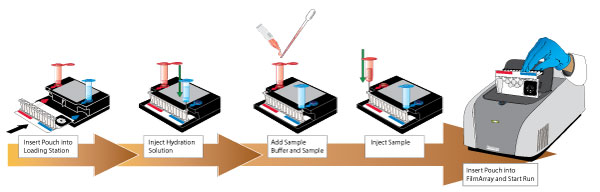
3) BIOFIRE Microfilmarray, a one hour test that detects the presence of the mecA gene
Resistance is confirmed by susceptibility testing with resistance to Oxacillin.
There are a large number of these proteins, and each bacterial prototype may have several of them. PBPs are involved in the last stages of peptidoglycan synthesis and the cross-linking of the cell wall for its shape and structure. This is essential for bacterial growth, survival, and cellular division and reproduction. Destruction of PBPs by the Beta-lactam antibiotics results in cell wall irregularities, elongation (can be seen in the Gram stain of patients on these antibiotics), and cell death. When the Beta-lactam antibiotic binds to this protein, the Beta-lactam amide bond is broken. This results in the formation of a covalent bond at the PBP's active site with the catalytic serine residue, and this bond is irreversible, thereby inactivating the protein permanently.
In cases of antibiotic resistance to Beta-lactams, such as in Methicillin-Resistant Staphylococcus Aureus, its PBP2A protein has a low affinity for penicillin and methicillin and the Beta-lactams, so the bond is not tight. This permits the protein to continue synthesizing the cell wall in the presence of the antibiotic and transpeptidase enzyme. This prevents the inhibiting of cell wall synthesis and renders the bacterium resistant to the drug(s). The presence of the PBP2A protein, and the presence of the mecA gene which encodes for it, enable this bacterium a resistant organism.
There are several rapid microbiology tests that detect the presence of the PBP2A protein or the mecA gene. These include the:
1) PBP2A rapid test, a quick 5 minute screen that detects the presence of this protein and presumptive MRSA
2) MRSA PCR (GenEXPERT), a one hour test that detects the presence of the mecA gene
3) BIOFIRE Microfilmarray, a one hour test that detects the presence of the mecA gene
Resistance is confirmed by susceptibility testing with resistance to Oxacillin.
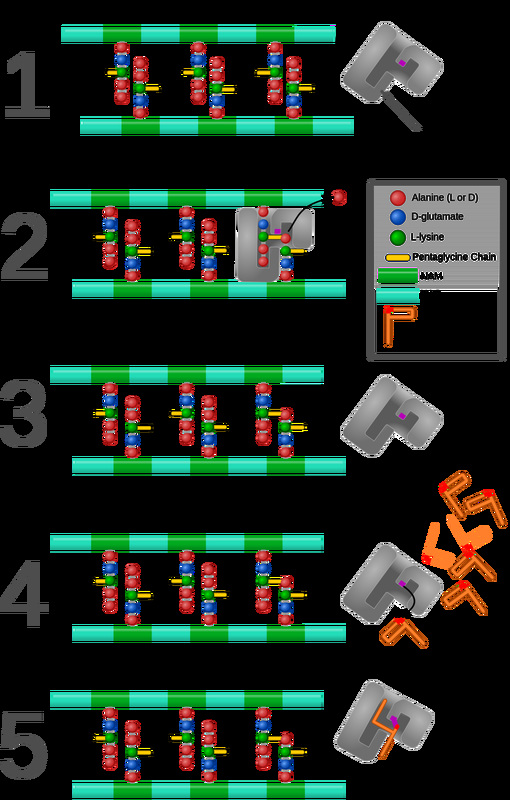
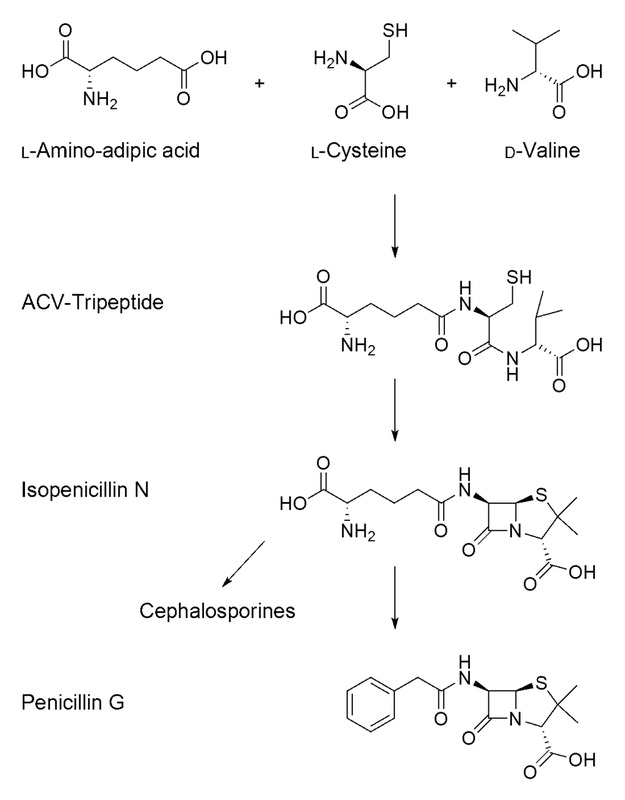
The building of peptidoglycan in the bacterial cell wall and the presence of PBP, which is responsible for catalyzing the cross-linking of the bacterial cell wall. In this scenario, penicillin, which has a structure similar to the PBP protein, is binding the PBP and results in its destruction, thereby permanently inhibiting cell wall synthesis and disrupting the cell, leading to death and lysis.